PI Steen Rasmussen, Harald Fellermann SDU
A simulation framework to analyze phase transitions of micelles to vesicles and gels of scpDNAs was developed. Relevant physical forces that need to be considered by such a simulation framework are (a) hydrophobic interactions that drive the aggregation of lipids and/or copolymer parts of scpDNAs, (b) hydrogen bonds that drive the Watson-Crick pairing of the nucleobase part of scpDNAs, and (c) electrostatic interactions that are envisioned to be used to control the overall aggregation behavior of scpDNAs. Additionally, the framework has to allow for chemical reactions needed e.g. for scpDNA ligation.
Because of the length and time scale of interest, dissipative particle dynamics (DPD) and extensions thereof are the simulation method of choice for such systems: hydrophobic interactions are well-addressed by DPD as hydrophobic self-assembly is the very motivation for the development of the method. Hydrogen bonds have been incorporated into DPD by partners of the ECCell project via incorporation of 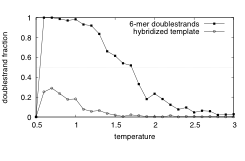 directional force fields into the method, where directions are either defined by covalent bonds [2] or dipole moments of single DPD beads [3]. Electrostatic inter-actions in DPD have been addressed in the literature by solving the Poisson equation [4] as well as the more conventional Edward summation [5]. Finally, chemical reactions have also been incorporated into DPD in earlier projects [6]
directional force fields into the method, where directions are either defined by covalent bonds [2] or dipole moments of single DPD beads [3]. Electrostatic inter-actions in DPD have been addressed in the literature by solving the Poisson equation [4] as well as the more conventional Edward summation [5]. Finally, chemical reactions have also been incorporated into DPD in earlier projects [6]
Figure 4: Hybridization of nucleic acids. Time fraction of two complementary nucleic acid 6-mers (upper curve), and one 6-mer with two complementary 3-mers (lower curve) at a lipid-water interface as a function of temperature
Using the DPD extensions of Ref. [2], we have studied the self-replication dynamics (consisting of hybridization, ligation, and dehybridization) of nucleic acids at oil-water interfaces. In order to achieve dehybridization of ligated nucleic acid strands, a simple mechanism has been developed to temper DPD systems. Fig. 4 shows an example for the melting behavior of ligated nucleic acid polymers and oligomers as a function of temperature.
[1] M. S. DeClue, P.-A. Monnard, J. Bailey, S.E. Maurer, G.E. Colin, H. Ziock, J. W. Boncella, S. and Rasmussen Nucleobase Mediated, Photocatalytic Vesicle Formation from an Ester Precursor Molecule. J. Am. Chem. Soc., 131, 931-933, 2009.
[2] H. Fellermann, S. Rasmussen, H.-J. Ziock, and R. Solé, Life-cycle of a minimal protocell – a dissipative particle dynamics (DPD) study, Artif. Life 13(4):319-345, 2007
[3] R.M. Füchslin, T. Maeke, and J.S. McCaskill, Spatially resolved simulations of membrane reactions and dynamics: Multipolar reaction DPD, Eur. Phys. J. E, DOI 10.1140/ep je/i2009-10482-x, 2009
[4] R.D. Groot, Electrostatic interactions in dissipative particle dynamics – simulation of polyelectrolytes and anionic surfactants, J. Chem. Phys. 118(24):11265-11277, 2003
[5] M. González-Melchor, E. Mayoral, M.E. Velázquez, and J. Alejandre,
Electrostatic interactions in dissipative particle dynamics using the Ewald sums, J. Chem. Phys. 125:224107, 2006
[6] H. Fellermann, Spatially resolved artificial chemistry, In: A. Adamatzky and M. Komosinski (eds.), Artificial Life Models in Software 2nd Edition, Springer London, 2009
