Itamar Willner
The Institute of Chemistry, The Hebrew University of Jerusalem
Objectives 2009-10
The objectives during this year of research included:
- The synthesis of Au nanoparticles (NPs) matrices on electrode surfaces for the programmed electrochemical uptake and release of substrates.
- The development of DNA-stimulated autocatalytic systems that enable DNA detection.
Electrochemically controlled uptake and release of substrates by Au NPs composites associated with electrodes
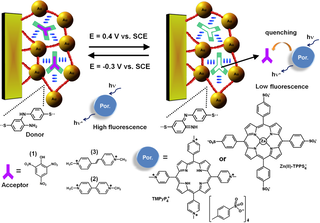
Figure 1. Photonic imaging of the electro-stimulated uptake and release of π-acceptor molecules using the imprinted bis-aniline-crosslinked Au NPs composite.
A method to electrosynthesize bis-aniline-bridged Au NPs composites on electrodes, by the electropolymerization of thioaniline-functionalized Au NPs, was developed. The electropolymerization of the Au NPs in the presence of π-acceptor substrates, such as picric acid (1), N,N-dimethyl-4,4'-bipyridinium (2) or N,N'-dimethyl-dipyridinium ethylene, (3), resulted in the imprinting of molecular recognition sites for the respective substrates into the Au NPs composite, Figure 1. As the bis-aniline bridging units reveal quasi-reversible redox properties the respective π-acceptor substrates, (1) – (3), bind to the imprinted sites via π donor-acceptor interactions with the bis-aniline bridging units. The electrochemical oxidation of the bis-aniline bridging units to the π-acceptor quinoid state, eliminates the affinity interactions between the substrates (1) – (3) and the bridging units, resulting in the release of the substrates from the Au NPs composite. By the cyclic reduction of the molecular bridging units to their π-donor bis-aniline state and the reverse oxidation of the bridging units to the quinoid state, the reversible uptake and release of the substrates to and from the Au NPs matrices was demonstrated. The activity of the Au NPs composites as "electrified sponges" was probed by following the fluorescence changes of protoporphyrin IX or Zn(II)-meso-tetraphenylsulfonate. Figure 2 exemplifies the fluorescence changes observed upon the uptake and release of (2).
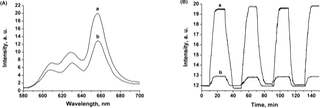
Figure 2. (A) Emission spectra corresponding to the Zn(II)-meso-tetraphenyl sulfonato porphyrin, Zn(II)-TPPS4-, 30 nM, in a 0.1 M HEPES buffer solution (pH=7.2), following the application of: (a) Reduction potential, E=-0.3 V vs. Ag QRE, for 10 minutes, and, (b) Oxidation potential, E=0.4 V vs. Ag QRE, for 10 minutes on the MV2+-imprinted bis-aniline-crosslinked Au NPs composite-modified Pt black-roughened Au electrode. (B) Switchable fluorescence intensity changes, at λ=660 nm, corresponding to Zn(II)-TPPS4-, 30 nM, in a 0.1 M HEPES buffer solution (pH=7.2), following the cyclic electro-stimulated uptake and release of MV2+ by: (a) the MV2+-imprinted bis-aniline-crosslinked Au NPs composite-modified Pt black-roughened Au electrode, (b) The non-imprinted bis-aniline-crosslinked Au NPs composite-modified Pt black-roughened Au electrode. In all the experiments the concentration of MV2+ was 2 nM and the excitation wavelength was λex=445.
The study has highlighted the following important conclusions:
- The imprinted sites bind effectively and specifically the respective imprinted substrates. The non-imprinted Au NPs matrices revealed low affinities for the substrates. As a result, the imprinted Au NPs matrices revealed high capacities for the association of the imprinted substrates.
- Enhanced binding capacities for the association of the imprinted substrates was achieved by the primary deposition of Pt black on the electrode, and the deposition of the imprinted Au NPs composites on the high-surface-area Pt support.
The detection of DNA by the isothermal autonomous replication of the analyte.
The development of isothermal autocatalytic systems that could replace the polymerase chain reaction (PCR) is a major goal in bioanalysis. Within the research efforts of the project, we developed an autocatalytic isothermal DNA detection platform using DNAzymes as biocatalytic labels, and fluorescence as readout signals. The sensing platform is depicted in Figure 3. The system consists of a subunit of the Mg2+-dependent DNAzyme, (I), and a hairpin structure (II) that includes in its stem region the complementary subunit Mg2+-dependent DNAzyme in a blocked configuration, and a recognition site for the analyte DNA in the loop region of the hairpin structure. The hairpin structure (III) includes in its single-stranded loop domain the ribonucleobase sequence that acts as a substrate for Mg2+-dependent DNAzyme, and its blocked stem region includes the sequence of the target DNA. The substrate hairpin (III) is modified by a fluorophore-quencher pair, and upon the DNAzyme-stimulated cleavage of III, the hairpin separates into the fragments, a process that triggers on the fluorescence of the fluorophore.
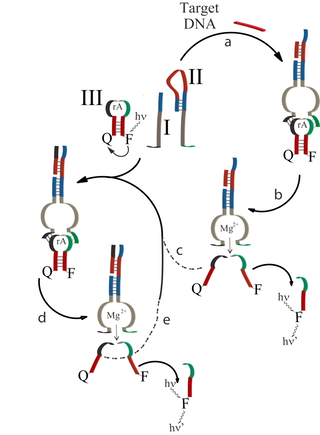
Accordingly, the opening of hairpin II by the target DNA activates the self-assembly of the DNAzyme subunits to the biocatalytic structure that cleaves the substrate III that releases the sequence analogous to the target. Thus, the autonomous autocatalytic generation of the target sequence by the detection of the target is initiated, resulting in the generation of fluorescence. Figure 4 depicts the time-dependent fluorescence intensities of the system upon the detection of variable concentrations of the target DNA (crosses indicate experimental data, whereas solid lines reflect the computer simulation of the experiments using the respective autocatalytic kinetic scheme).
Figure 3. Schematic representation of the analyte-induced DNAzyme assembly and the sensing process: (a) target recognition and assembly of the DNAzyme, (b) cleavage of the substrate 5 and release of the activator unit 6, (c-e) autocatalytic, catabolic generation cycle triggered by the activator-unit-induced assembly of the DNAzyme.
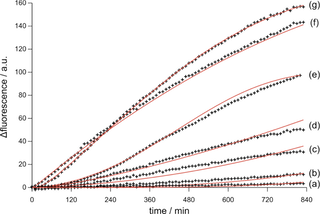
Figure 4. Time-dependent fluorescence increase (cross dots) and non-linear regression of the autocatalytic nucleic acid sensing with target concentrations of: (a) 1x10-12 M, (b) 1x10-11 M, (c) 1x10-10 M, (d) 1x10-9 M, (e) 1x10-8 M, (f) 1x10-7 M and (g) 1x10-6 M.
The major conclusions of the study are:
• An autocatalytic, isothermal, DNA detection scheme of DNA was developed. The detection limit for analyzing the target DNA was estimated to be 5 x 10-13 M.
• The simulation of the experimental results suggests that the activity of the Mg2+-dependent DNAzyme is low, and this limits the autocatalytic process. The use of DNAzymes of higher activities could improve the performance of the system.
Publications
1. M. Frasconi, R. Tel-Vered, J. Elbaz and I. Willner
Electrochemically Stimulated pH Changes: A Route to Control Chemical Reactivity.
J. Am. Chem. Soc., 132, 2029-2036 (2010).
2. M. Frasconi, R. Tel-Vered, M. Riskin and I. Willner
Electrified Selective "Sponges" Made of Au Nanoparticles.
J. Am. Chem. Soc., in press (2010)
3. F. Wang, J. Elbaz, C. Teller and I. Willner
Amplified Detection of DNA through the Autocatalytic, Catabolic Generation of the Analyte by a DNAzyme. Submitted for publication.
Future Plans
• The design of functional Au NPs composites acting as light-activated "sponges" for the uptake/release of substrates.
• The application of alternative DNAzymes for the amplified detection of DNA using isothermal autocatalytic processes.
• Designing pH-switchable DNAzymes.
