Fakultät für
Chemie
und Biochemie
Biochemie II
Biomolekulare NMR-Spektroskopie
Prof. Dr. Raphael Stoll
Gebäude NC 5/171
Universitätsstraße 150
D-44780 Bochum
Tel.: +49 234 32-25466
Fax: +49 234 32-05466
E-Mail: bionmr@rub.de
Projekte
Bulky cargos like procollagens, apolipoproteins, and mucins exceed the size of conventional COPII vesicles. During evolution a process emerged in metazoans, predominantly governed by the TANGO1 protein family, that organizes cargo at the exit sites of the endoplasmic reticulum and facilitates export by the formation of tunnel-like connections between the ER and Golgi. Hitherto, cargo-recognition appeared to be mediated by an SH3-like domain. Based on structural and dynamic data as well as interaction studies from NMR spectroscopy and microscale thermophoresis presented here, we show that the luminal cargo-recognition domain of TANGO1 adopts a new functional fold for which we suggest the term MOTH (MIA, Otoraplin, TALI/TANGO1 homology) domain. These MOTH domains, as well as an evolutionary intermediate found in invertebrates, constitute a distinct domain family that emerged from SH3 domains and acquired the ability to bind collagen.
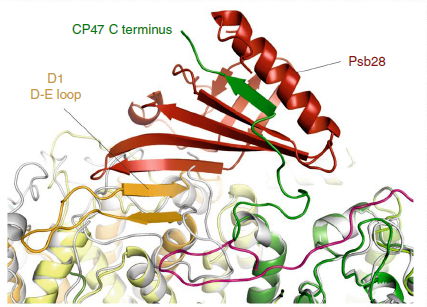
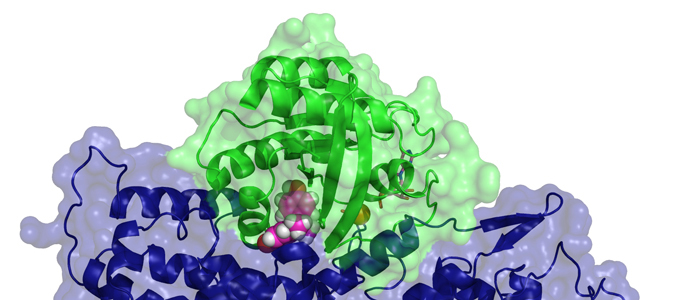
Biogenesis of photosystem II (PSII), nature's water-splitting catalyst, is assisted by auxiliary proteins that form transient complexes with PSII components to facilitate stepwise assembly events. Using cryo-electron microscopy, we solved the structure of such a PSII assembly intermediate from Thermosynechococcus elongatus at 2.94 Å resolution. It contains three assembly factors (Psb27, Psb28 and Psb34) and provides detailed insights into their molecular function. Binding of Psb28 induces large conformational changes at the PSII acceptor side, which distort the binding pocket of the mobile quinone (QB) and replace the bicarbonate ligand of non-haem iron with glutamate, a structural motif found in reaction centres of non-oxygenic photosynthetic bacteria. These results reveal mechanisms that protect PSII from damage during biogenesis until water splitting is activated. Our structure further demonstrates how the PSII active site is prepared for the incorporation of the Mn4CaO5 cluster, which performs the unique water-splitting reaction.
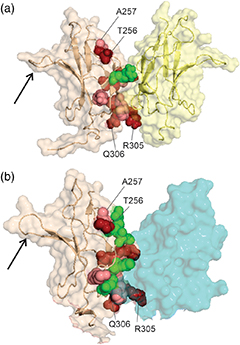 "The nuclear factor of kappa light polypeptide gene enhancer in B-cells (NFκB) transcription factors play a critical role in human immune response. The family includes homodimers and heterodimers of five component proteins, which mediate different transcriptional responses and bind preferentially to different DNA sequences. Crystal structures of DNA complexes show that the dimers of the Rel-homology regions are structurally very similar. Differing DNA sequence preference together with structural similarity suggests that the dimers may differ in their dynamics. In this study, we present the first near-complete 15 N, 13 Cα/β , and HN backbone resonance assignments of two dimers of the dimerization domain (DD) of the NFκB1 (p50) protein (residues 241-351): the homodimer of two p50 domains and a heterodimer of the p50 DD with the p65 DD. As expected, the two dimers behave very similarly, with chemical shift differences between them largely concentrated in the dimer interface and attributable to specific differences in the amino acid sequences of p50 and p65. A comparison of the picosecond-nanosecond dynamics of the homo- and heterodimers also shows that the environment of p50 is similar, with an overall slightly reduced correlation time for the homodimer compared to the heterodimer, consistent with its slightly smaller molecular weight. These results demonstrate that NMR spectroscopy can be used to explore subtle changes in structure and dynamics that have the potential to give insights into differences in specificity that can be exploited in the design of new therapeutic agents.
"The nuclear factor of kappa light polypeptide gene enhancer in B-cells (NFκB) transcription factors play a critical role in human immune response. The family includes homodimers and heterodimers of five component proteins, which mediate different transcriptional responses and bind preferentially to different DNA sequences. Crystal structures of DNA complexes show that the dimers of the Rel-homology regions are structurally very similar. Differing DNA sequence preference together with structural similarity suggests that the dimers may differ in their dynamics. In this study, we present the first near-complete 15 N, 13 Cα/β , and HN backbone resonance assignments of two dimers of the dimerization domain (DD) of the NFκB1 (p50) protein (residues 241-351): the homodimer of two p50 domains and a heterodimer of the p50 DD with the p65 DD. As expected, the two dimers behave very similarly, with chemical shift differences between them largely concentrated in the dimer interface and attributable to specific differences in the amino acid sequences of p50 and p65. A comparison of the picosecond-nanosecond dynamics of the homo- and heterodimers also shows that the environment of p50 is similar, with an overall slightly reduced correlation time for the homodimer compared to the heterodimer, consistent with its slightly smaller molecular weight. These results demonstrate that NMR spectroscopy can be used to explore subtle changes in structure and dynamics that have the potential to give insights into differences in specificity that can be exploited in the design of new therapeutic agents.
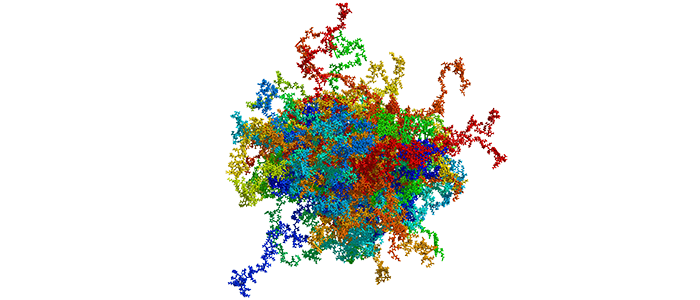
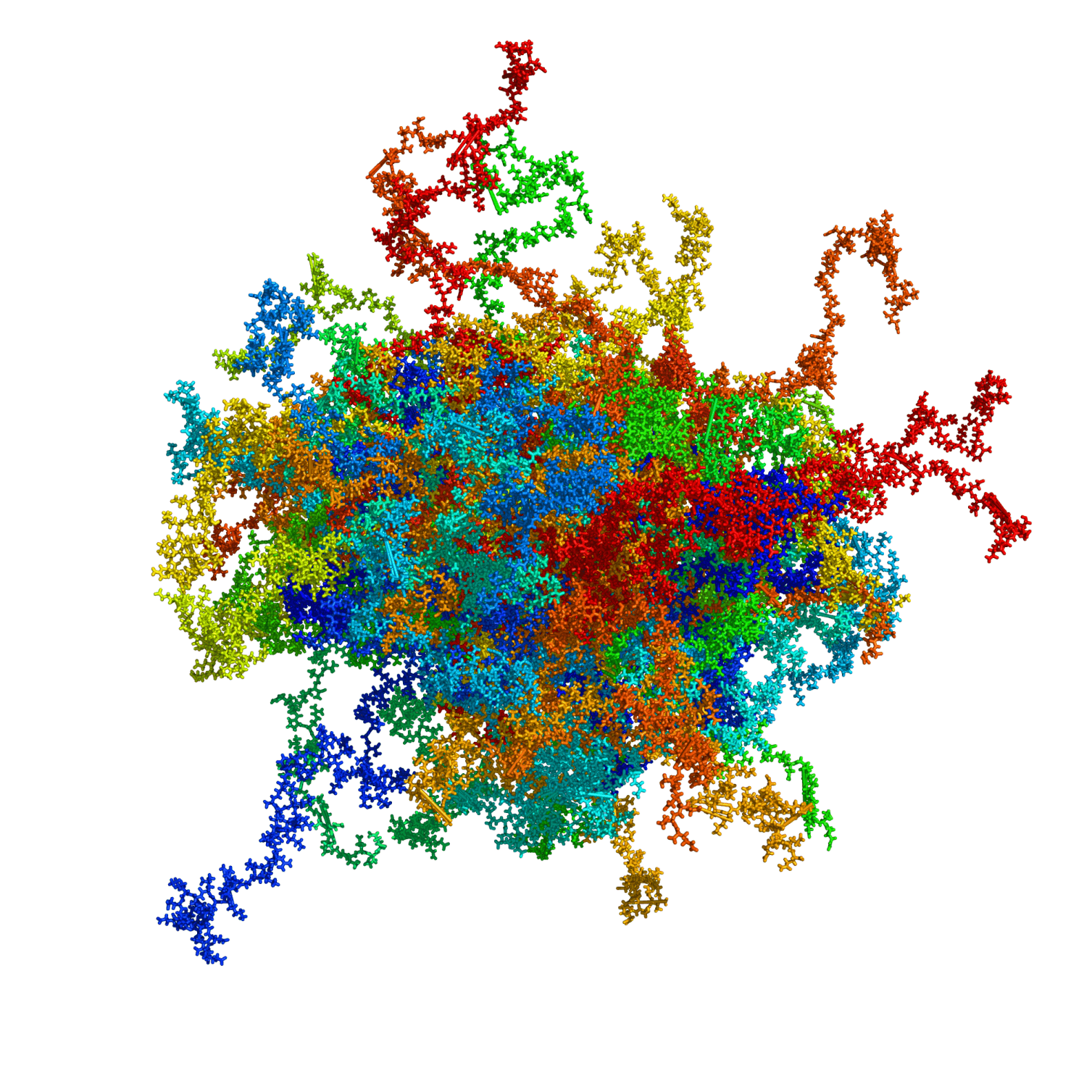 High Mobility Group Protein A1a (HMGA1a) is a highly abundant nuclear protein, which plays a crucial role during embryogenesis, cell differentiation, and neoplasia. Here, we present the first ever NMR-based structural ensemble of full length HMGA1a. Our results show that the protein is not completely random coil but adopts a compact structure consisting of transient long-range contacts, which is regulated by post-translational phosphorylation. The CK2-, cdc2- and cdc2/CK2-phosphorylated forms of HMGA1a each exhibit a different binding affinity towards the PRD2 element of the NFκB promoter. Our study identifies connected regions between phosphorylation sites in the wildtype ensemble that change considerably upon phosphorylation, indicating that these posttranslational modifications sites are part of an electrostatic contact network that alters the structural ensemble by shifting the conformational equilibrium. Moreover, ITC data reveal that the CK2-phosphorylated HMGA1a exhibits a different DNA promoter binding affinity for the PRD2 element. Furthermore, we present the first structural model for AT-hook 1 of HMGA1a that can adopt a transient α-helical structure, which might serve as an additional regulatory mechanism in HMAG1a. Our findings will help to develop new therapeutic strategies against HMGA1a-associated cancers by taking posttranslational modifications into consideration.
High Mobility Group Protein A1a (HMGA1a) is a highly abundant nuclear protein, which plays a crucial role during embryogenesis, cell differentiation, and neoplasia. Here, we present the first ever NMR-based structural ensemble of full length HMGA1a. Our results show that the protein is not completely random coil but adopts a compact structure consisting of transient long-range contacts, which is regulated by post-translational phosphorylation. The CK2-, cdc2- and cdc2/CK2-phosphorylated forms of HMGA1a each exhibit a different binding affinity towards the PRD2 element of the NFκB promoter. Our study identifies connected regions between phosphorylation sites in the wildtype ensemble that change considerably upon phosphorylation, indicating that these posttranslational modifications sites are part of an electrostatic contact network that alters the structural ensemble by shifting the conformational equilibrium. Moreover, ITC data reveal that the CK2-phosphorylated HMGA1a exhibits a different DNA promoter binding affinity for the PRD2 element. Furthermore, we present the first structural model for AT-hook 1 of HMGA1a that can adopt a transient α-helical structure, which might serve as an additional regulatory mechanism in HMAG1a. Our findings will help to develop new therapeutic strategies against HMGA1a-associated cancers by taking posttranslational modifications into consideration.
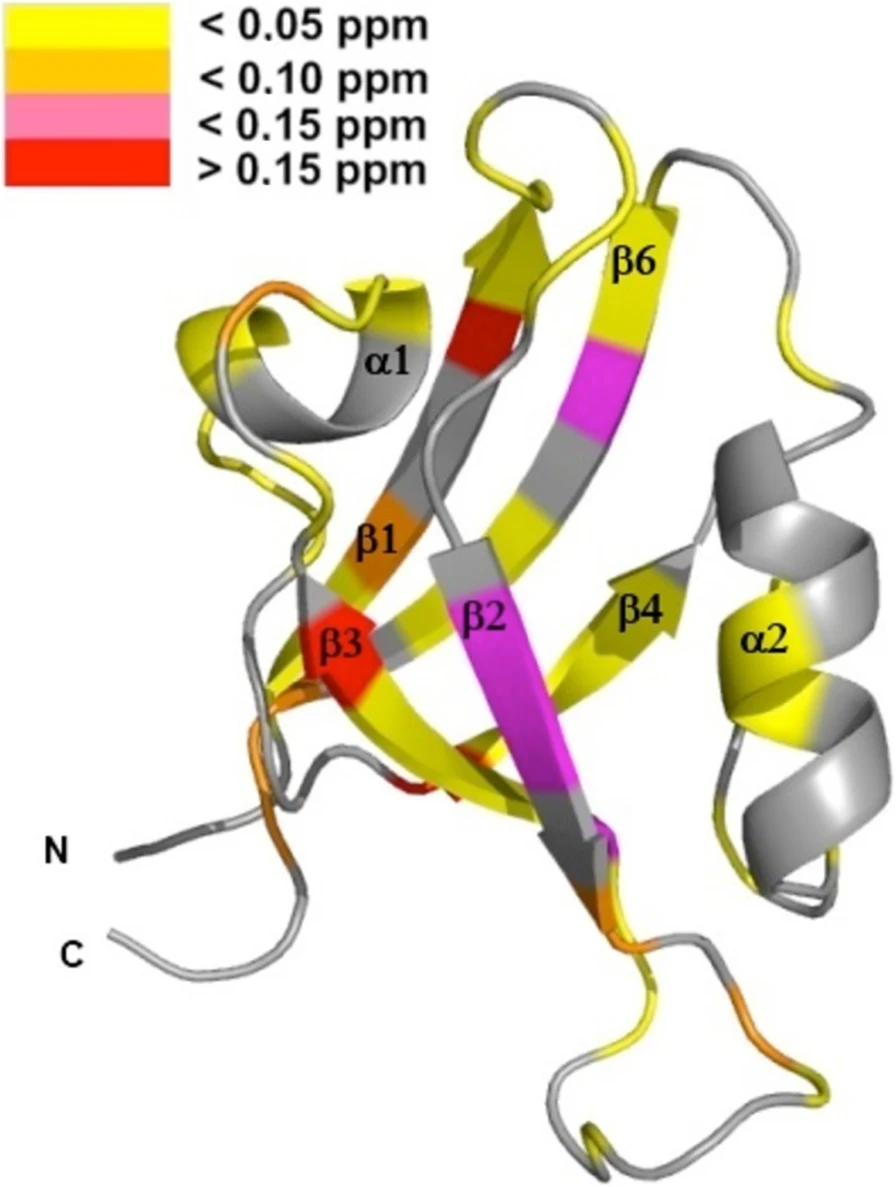 Protein tyrosine phosphatase PTPN13, also known as PTP-BL in mice, is a large multi-domain non-transmembrane scaffolding protein with a molecular mass of 270 kDa. It is involved in the regulation of several cellular processes such as cytokinesis and actin-cytoskeletal rearrangement. The modular structure of PTPN13 consists of an N-terminal KIND domain, a FERM domain, and five PDZ domains, followed by a C-terminal protein tyrosine phosphatase domain. PDZ domains are among the most abundant protein modules and they play a crucial role in signal transduction of protein networks.
Protein tyrosine phosphatase PTPN13, also known as PTP-BL in mice, is a large multi-domain non-transmembrane scaffolding protein with a molecular mass of 270 kDa. It is involved in the regulation of several cellular processes such as cytokinesis and actin-cytoskeletal rearrangement. The modular structure of PTPN13 consists of an N-terminal KIND domain, a FERM domain, and five PDZ domains, followed by a C-terminal protein tyrosine phosphatase domain. PDZ domains are among the most abundant protein modules and they play a crucial role in signal transduction of protein networks.
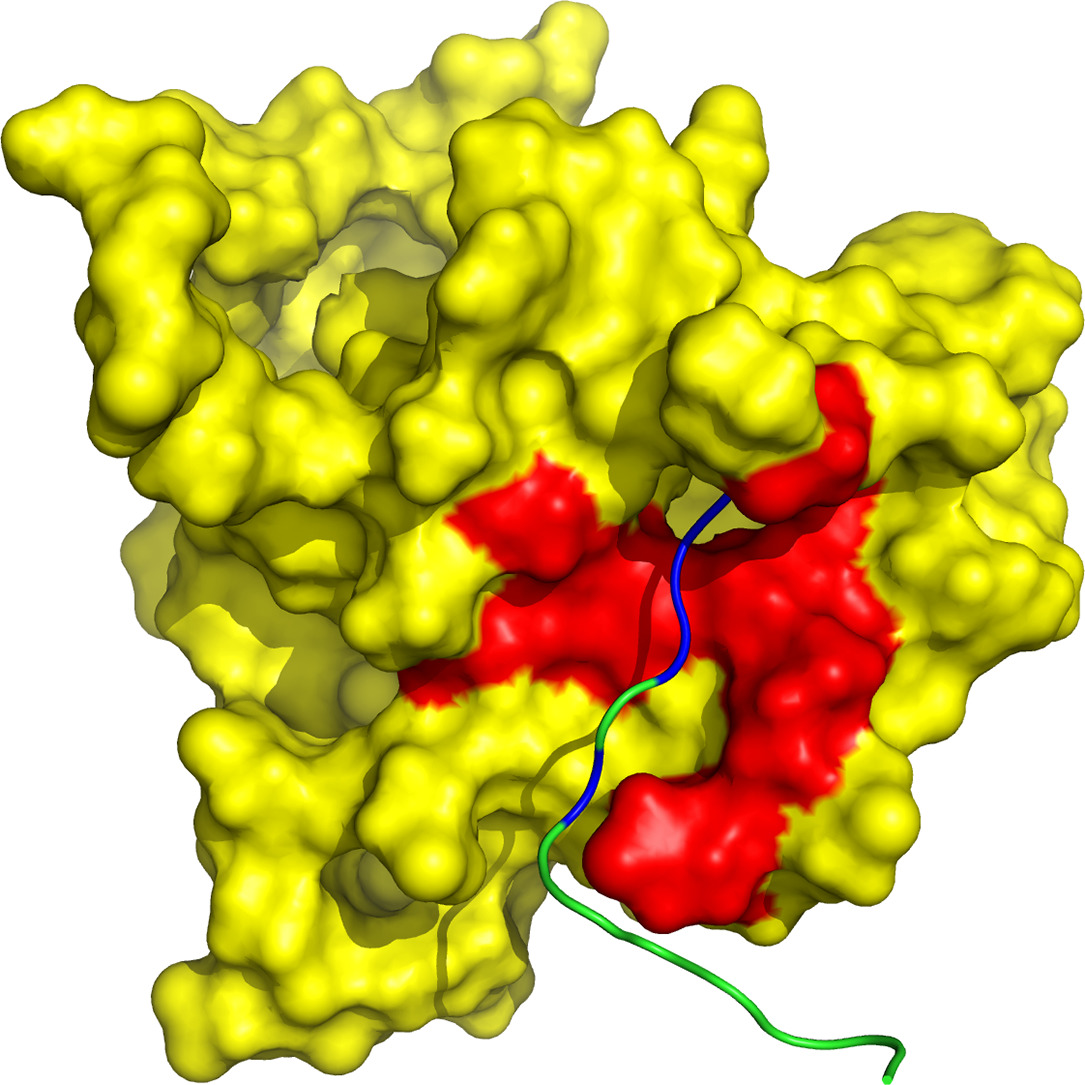 Protein tyrosine phosphatase PTPN13, also known as PTP-BL in mice, represents a large multi-domain non-transmembrane scaffolding protein that contains five consecutive PDZ domains. Here, we report the solution structures of the extended murine PTPN13 PDZ3 domain in its apo form and in complex with its physiological ligand, the carboxy-terminus of protein kinase C-related kinase-2 (PRK2), determined by multidimensional NMR spectroscopy. Both in its ligand-free state and when complexed to PRK2, PDZ3 of PTPN13 adopts the classical compact, globular D/E fold. PDZ3 of PTPN13 binds five carboxy-terminal amino acids of PRK2 via a groove located between the EB-strand and the DB-helix. The PRK2 peptide resides in the canonical PDZ3 binding cleft in an elongated manner and the amino acid side chains in position P0 and P-2, cysteine and aspartate, of the ligand face the groove between EB-strand and DB-helix, whereas the PRK2 side chains of tryptophan and alanine located in position P-1 and P-3 point away from the binding cleft. These structures are rare examples of selective class III ligand recognition by a PDZ domain and now provide a basis for the detailed structural investigation of the promiscuous interaction between the PDZ domains of PTPN13 and their ligands. They will also lead to a better understanding of the proposed scaffolding function of these domains in multi-protein complexes assembled by PTPN13 and could ultimately contribute to low molecular weight antagonists that might even act on the PRK2 signaling pathway to modulate rearrangements of the actin cytoskeleton.
Protein tyrosine phosphatase PTPN13, also known as PTP-BL in mice, represents a large multi-domain non-transmembrane scaffolding protein that contains five consecutive PDZ domains. Here, we report the solution structures of the extended murine PTPN13 PDZ3 domain in its apo form and in complex with its physiological ligand, the carboxy-terminus of protein kinase C-related kinase-2 (PRK2), determined by multidimensional NMR spectroscopy. Both in its ligand-free state and when complexed to PRK2, PDZ3 of PTPN13 adopts the classical compact, globular D/E fold. PDZ3 of PTPN13 binds five carboxy-terminal amino acids of PRK2 via a groove located between the EB-strand and the DB-helix. The PRK2 peptide resides in the canonical PDZ3 binding cleft in an elongated manner and the amino acid side chains in position P0 and P-2, cysteine and aspartate, of the ligand face the groove between EB-strand and DB-helix, whereas the PRK2 side chains of tryptophan and alanine located in position P-1 and P-3 point away from the binding cleft. These structures are rare examples of selective class III ligand recognition by a PDZ domain and now provide a basis for the detailed structural investigation of the promiscuous interaction between the PDZ domains of PTPN13 and their ligands. They will also lead to a better understanding of the proposed scaffolding function of these domains in multi-protein complexes assembled by PTPN13 and could ultimately contribute to low molecular weight antagonists that might even act on the PRK2 signaling pathway to modulate rearrangements of the actin cytoskeleton.
The protein family of small GTPases controls cellular processes by acting as a binary switch between an active and an inactive state. The most prominent family members are H-Ras, N-Ras, and K-Ras isoforms, which are highly related and frequently mutated in cancer. Bisphenols are widespread in modern life because of their industrial application as plasticisers. Bisphenol A (BPA) is the best-known member and has gained significant scientific as well as public attention as an endocrine disrupting chemical, a fact that eventually led to its replacement. However, compounds used to replace BPA still contain the molecular scaffold of bisphenols. Here we show that BPA, BPAF, BPB, BPE, BPF, and an amine-substituted BPAF-derivate all interact with all GDP-bound Ras-Isoforms through binding to a common site on these proteins. NMR-, SOScat-, and GDI- assay-based data revealed a new bisphenol-induced, allosterically activated GDP-bound Ras conformation that define these plasticisers as Ras allosteric agonists.
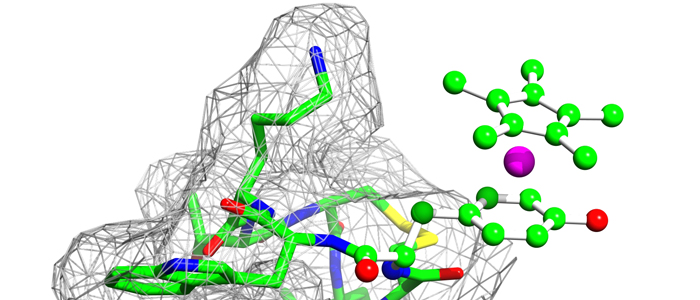
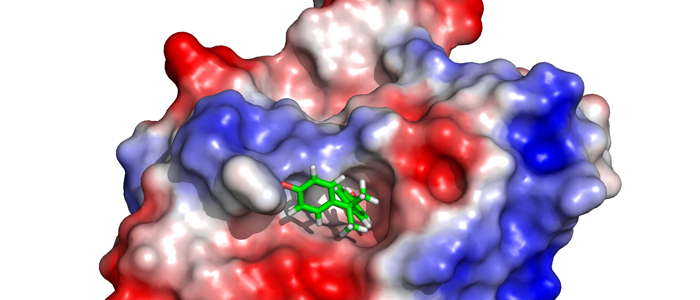
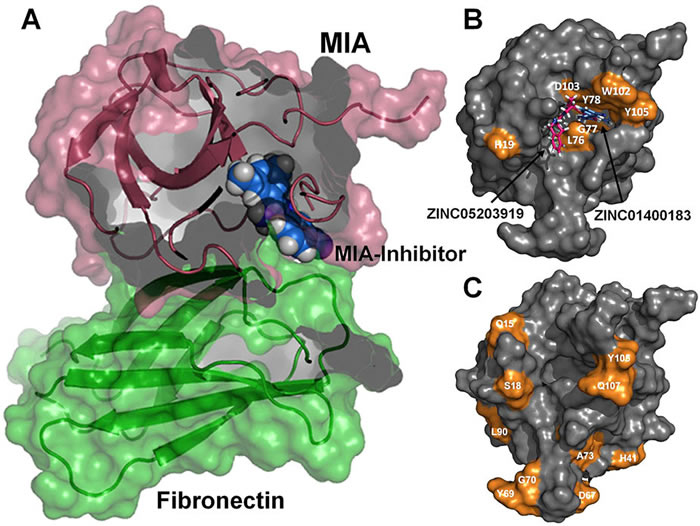
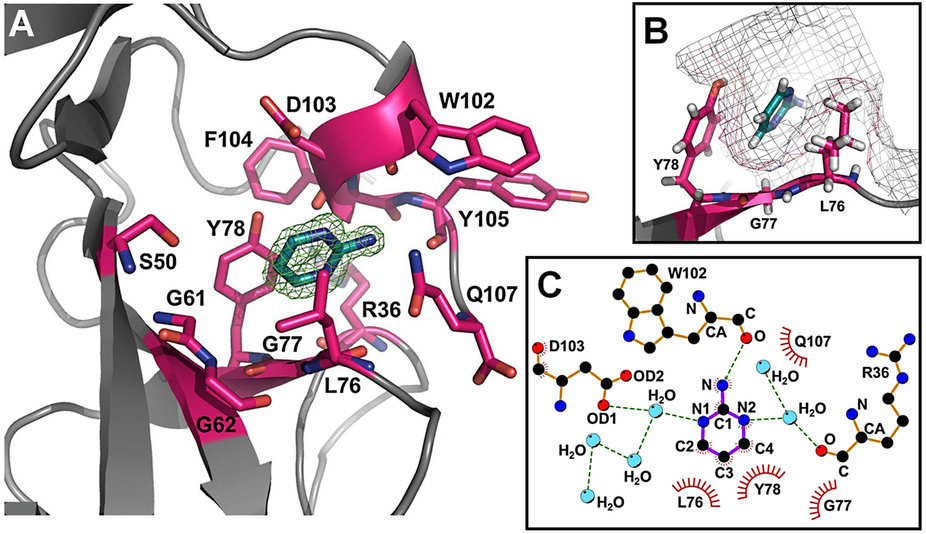 Melanoma inhibitory activity (MIA), an extracellular protein highly expressed by malignant melanoma cells, plays an important functional role in melanoma development, progression, and metastasis. After its secretion, MIA directly interacts with extracellular matrix proteins, such as fibronectin (FN). By this mechanism, MIA actively facilitates focal cell detachment from surrounding structures and strongly promotes tumour cell invasion and migration. Hence, the molecular understanding of MIA’s function provides a promising target for the development of new strategies in malignant melanoma therapy. Here, we describe for the first time the discovery of small molecules that are able to disrupt the MIA-FN complex by selectively binding to a new druggable pocket, which we could identify on MIA by structural analysis and fragment-based screening. Our findings may inspire novel drug discovery efforts aiming at a therapeutically effective treatment of melanoma by targeting MIA.
Melanoma inhibitory activity (MIA), an extracellular protein highly expressed by malignant melanoma cells, plays an important functional role in melanoma development, progression, and metastasis. After its secretion, MIA directly interacts with extracellular matrix proteins, such as fibronectin (FN). By this mechanism, MIA actively facilitates focal cell detachment from surrounding structures and strongly promotes tumour cell invasion and migration. Hence, the molecular understanding of MIA’s function provides a promising target for the development of new strategies in malignant melanoma therapy. Here, we describe for the first time the discovery of small molecules that are able to disrupt the MIA-FN complex by selectively binding to a new druggable pocket, which we could identify on MIA by structural analysis and fragment-based screening. Our findings may inspire novel drug discovery efforts aiming at a therapeutically effective treatment of melanoma by targeting MIA.
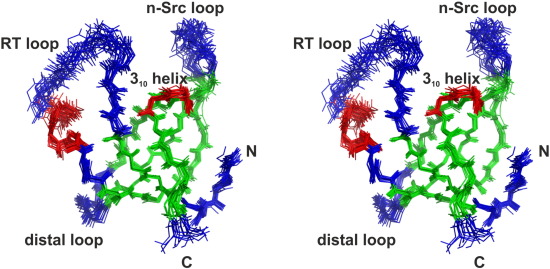 PetP is a peripheral subunit of the cytochrome b6f complex (b6f) present in both, cyanobacteria and red algae. It
is bound to the cytoplasmic surface of this membrane protein complex where it greatly affects the efficiency of
the linear photosynthetic electron flow although it is not directly involved in the electron transfer reactions.
Despite the crystal structures of the b6f core complex, structural information for the transient regulatory b6f subunits is still missing. Here we present the first structure of PetP at atomic resolution as determined by solution
NMR. The protein adopts an SH3 fold, which is a common protein motif in eukaryotes but comparatively rare
in prokaryotes. The structure of PetP enabled the identification of the potential interaction site for b6f binding
by conservation mapping. The interaction surface is mainly formed by two large loop regions and one short
310 helix which also exhibit an increased flexibility as indicated by heteronuclear steady-state {1H}-15N NOE
and randomcoil index parameters. The properties of this potential b6f binding site greatly differ fromthe canonical
peptide binding site which is highly conserved in eukaryotic SH3 domains. Interestingly, three other proteins
of the photosynthetic electron transport chain share this SH3 fold with PetP: NdhS of the photosynthetic NADH
dehydrogenase-like complex (NDH-1), PsaE of the photosystem 1 and subunit α of the ferredoxin–thioredoxin
reductase have, similar to PetP, a great impact on the photosynthetic electron transport. Finally, a model is presented to illustrate how SH3 domains modulate the photosynthetic electron transport processes in
cyanobacteria.
PetP is a peripheral subunit of the cytochrome b6f complex (b6f) present in both, cyanobacteria and red algae. It
is bound to the cytoplasmic surface of this membrane protein complex where it greatly affects the efficiency of
the linear photosynthetic electron flow although it is not directly involved in the electron transfer reactions.
Despite the crystal structures of the b6f core complex, structural information for the transient regulatory b6f subunits is still missing. Here we present the first structure of PetP at atomic resolution as determined by solution
NMR. The protein adopts an SH3 fold, which is a common protein motif in eukaryotes but comparatively rare
in prokaryotes. The structure of PetP enabled the identification of the potential interaction site for b6f binding
by conservation mapping. The interaction surface is mainly formed by two large loop regions and one short
310 helix which also exhibit an increased flexibility as indicated by heteronuclear steady-state {1H}-15N NOE
and randomcoil index parameters. The properties of this potential b6f binding site greatly differ fromthe canonical
peptide binding site which is highly conserved in eukaryotic SH3 domains. Interestingly, three other proteins
of the photosynthetic electron transport chain share this SH3 fold with PetP: NdhS of the photosynthetic NADH
dehydrogenase-like complex (NDH-1), PsaE of the photosystem 1 and subunit α of the ferredoxin–thioredoxin
reductase have, similar to PetP, a great impact on the photosynthetic electron transport. Finally, a model is presented to illustrate how SH3 domains modulate the photosynthetic electron transport processes in
cyanobacteria.
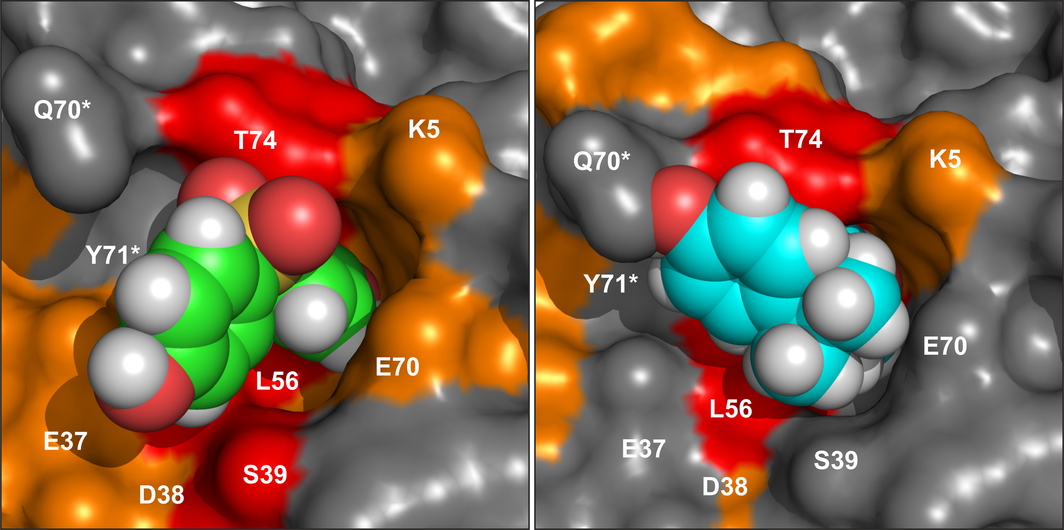 K-Ras4B is a small GTPase that belongs to the Ras superfamily of guanine nucleotide-binding proteins. GTPases function as molecular switches in cells and are key players in intracellular signalling. Ras has been identified as an oncogene and is mutated in more than 20% of human cancers. Here, we report that Bisphenol S binds into a binding pocket of K-Ras4B previously identified for various low molecular weight compounds. Our results advocate for more comprehensive safety studies on the toxicity of Bisphenol S, as it is frequently used for Bisphenol A-free food containers.
K-Ras4B is a small GTPase that belongs to the Ras superfamily of guanine nucleotide-binding proteins. GTPases function as molecular switches in cells and are key players in intracellular signalling. Ras has been identified as an oncogene and is mutated in more than 20% of human cancers. Here, we report that Bisphenol S binds into a binding pocket of K-Ras4B previously identified for various low molecular weight compounds. Our results advocate for more comprehensive safety studies on the toxicity of Bisphenol S, as it is frequently used for Bisphenol A-free food containers.
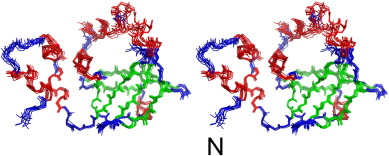 The cyanobacterial multi-subunit membrane protein complex NDH-1 is both structurally and functionally related to Complex I of eubacteria and mitochondria. In addition to functions in respiration and cyclic electron transfer around photosystem I (PSI), the cyanobacterial NDH-1 complex is involved in a unique mechanism for inorganic carbon concentration. Although the crystal structures of the similar respiratory Complex I from Thermus thermophilus and Escherichia coli are known, atomic structural information is not available for the cyanobacterial NDH-1 complex yet. In particular, the structures of those subunits that are not homologous to Complex I will help to understand their distinct functions. The 15.7kDa protein CupS is a small soluble subunit of the complex variant NDH-1MS, which is thought to play a role in CO2 conversion. Here, we present the NMR structure of CupS from Thermosynechococcus elongatus, which is the very first structure of a specific cyanobacterial NDH-1 complex subunit. CupS shares a structural similarity with members of the Fasciclin protein superfamily. The structural comparison to Fasciclin type proteins based on known NMR structures and protein sequences of human TGFBIp, MPB70 from Mycobacterium bovis, and Fdp from Rhodobacter sphaeroides, together with a virtual docking model of CupS and NdhF3, provide first insight into the specific binding of CupS to the NDH-1MS complex at atomic resolution.
The cyanobacterial multi-subunit membrane protein complex NDH-1 is both structurally and functionally related to Complex I of eubacteria and mitochondria. In addition to functions in respiration and cyclic electron transfer around photosystem I (PSI), the cyanobacterial NDH-1 complex is involved in a unique mechanism for inorganic carbon concentration. Although the crystal structures of the similar respiratory Complex I from Thermus thermophilus and Escherichia coli are known, atomic structural information is not available for the cyanobacterial NDH-1 complex yet. In particular, the structures of those subunits that are not homologous to Complex I will help to understand their distinct functions. The 15.7kDa protein CupS is a small soluble subunit of the complex variant NDH-1MS, which is thought to play a role in CO2 conversion. Here, we present the NMR structure of CupS from Thermosynechococcus elongatus, which is the very first structure of a specific cyanobacterial NDH-1 complex subunit. CupS shares a structural similarity with members of the Fasciclin protein superfamily. The structural comparison to Fasciclin type proteins based on known NMR structures and protein sequences of human TGFBIp, MPB70 from Mycobacterium bovis, and Fdp from Rhodobacter sphaeroides, together with a virtual docking model of CupS and NdhF3, provide first insight into the specific binding of CupS to the NDH-1MS complex at atomic resolution.
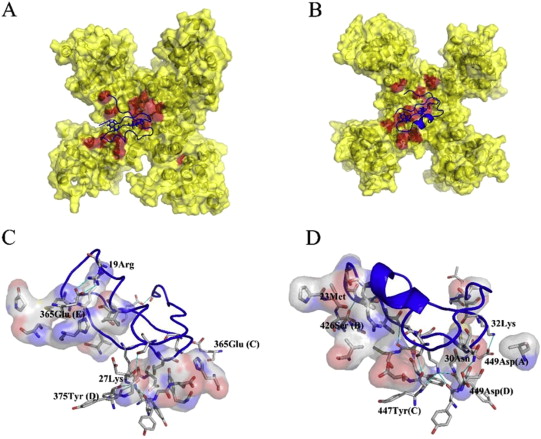 Animal venoms, such as those from scorpions, are a potent source for new pharmacological substances. In this study we have determined the structure of the α-KTx3.8 (named as Bs6) scorpion toxin by multidimensional 1H homonuclear NMR spectroscopy and investigated its function by molecular dynamics (MD) simulations and electrophysiological measurements. Bs6 is a potent inhibitor of the Kv1.3 channel which plays an important role during the activation and proliferation of memory T-cells (TEM), which play an important role in autoimmune diseases.
Therefore, it could be an interesting target for treatment of autoimmune diseases. In this study, Bs6 was synthesised by solid phase synthesis and its three-dimensional (3D) structure has been determined. To gain a deeper insight into the interaction of Bs6 with different potassium channels like hKv1.1 and hKv1.3, the protein-protein complex was modelled based on known toxin-channel structures and tested for stability in MD simulations using GROMACS. The toxin-channel interaction was further analysed by electrophysiological measurements of different potassium channels like hKv1.3 and hKv7.1. As potassium channel inhibitors could play an important role to overcome autoimmune diseases like multiple sclerosis and type-1 diabetes mellitus, our data contributes to the understanding of the molecular mechanism of action and will ultimately help to develop new potent inhibitors in future.
Animal venoms, such as those from scorpions, are a potent source for new pharmacological substances. In this study we have determined the structure of the α-KTx3.8 (named as Bs6) scorpion toxin by multidimensional 1H homonuclear NMR spectroscopy and investigated its function by molecular dynamics (MD) simulations and electrophysiological measurements. Bs6 is a potent inhibitor of the Kv1.3 channel which plays an important role during the activation and proliferation of memory T-cells (TEM), which play an important role in autoimmune diseases.
Therefore, it could be an interesting target for treatment of autoimmune diseases. In this study, Bs6 was synthesised by solid phase synthesis and its three-dimensional (3D) structure has been determined. To gain a deeper insight into the interaction of Bs6 with different potassium channels like hKv1.1 and hKv1.3, the protein-protein complex was modelled based on known toxin-channel structures and tested for stability in MD simulations using GROMACS. The toxin-channel interaction was further analysed by electrophysiological measurements of different potassium channels like hKv1.3 and hKv7.1. As potassium channel inhibitors could play an important role to overcome autoimmune diseases like multiple sclerosis and type-1 diabetes mellitus, our data contributes to the understanding of the molecular mechanism of action and will ultimately help to develop new potent inhibitors in future.
Nucleotide Exchange: Implications for GTPase-Selective Antagonists.
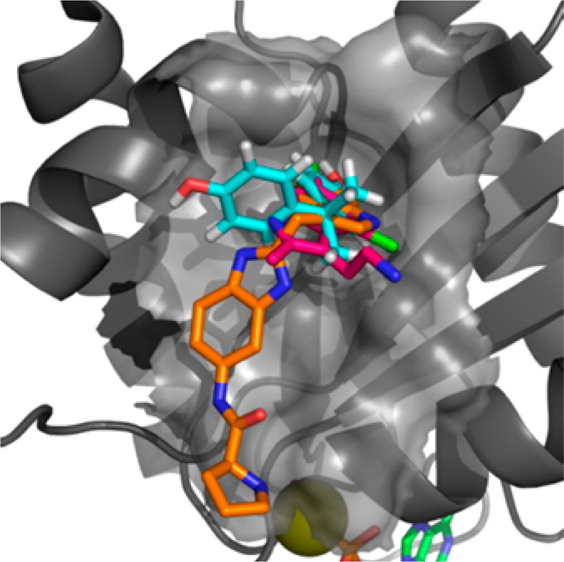 We show here for the first time that bisphenol A (10) has the capacity to interact directly with K-Ras and that Rheb weakly binds to bisphenol A (10) and 4,4′-biphenol derivatives. We have characterized these interactions at atomic resolution suggesting that these compounds sterically interfere with the Sos-mediated nucleotide exchange in H- and K-Ras. We show that 4,4′-biphenol (5) selectively inhibits Rheb signaling and induces cell death suggesting that this compound might be a novel candidate for treatment of tuberous sclerosis-mediated tumor growth. Our results propose a new mode of action for bisphenol A (10) that advocates a reduced exposure to this compound in our environment. Our data may lay the foundation for the future design of GTPaseselective antagonists with higher affinity to benefit of the treatment of cancer because KRas inhibition is regarded to be a promising strategy with a potential therapeutic window for targeting Sos in Ras-driven tumors.
We show here for the first time that bisphenol A (10) has the capacity to interact directly with K-Ras and that Rheb weakly binds to bisphenol A (10) and 4,4′-biphenol derivatives. We have characterized these interactions at atomic resolution suggesting that these compounds sterically interfere with the Sos-mediated nucleotide exchange in H- and K-Ras. We show that 4,4′-biphenol (5) selectively inhibits Rheb signaling and induces cell death suggesting that this compound might be a novel candidate for treatment of tuberous sclerosis-mediated tumor growth. Our results propose a new mode of action for bisphenol A (10) that advocates a reduced exposure to this compound in our environment. Our data may lay the foundation for the future design of GTPaseselective antagonists with higher affinity to benefit of the treatment of cancer because KRas inhibition is regarded to be a promising strategy with a potential therapeutic window for targeting Sos in Ras-driven tumors.
 Binding
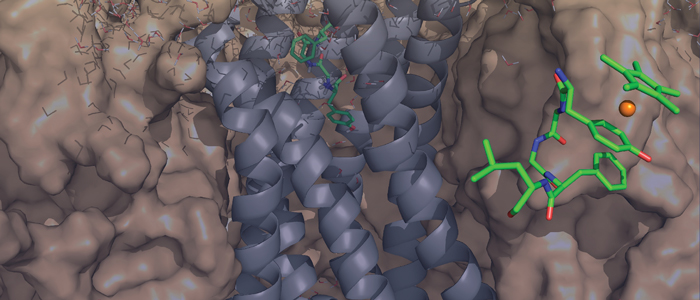
of Leu-enkephalin and [RhIII(η5-Cp*)(η6-Tyr1)]Leu-enkephalin
to the recently published crystal structures of the μ- and δ-opioid
receptor is studied. Docking of free Leu-enkephalin reveals two
preferred conformations, one of which suggests an alternative binding
site for the tyrosine residue. Furthermore, the three-dimensional
solution structure of [RhIII(η5-Cp*)(η6-Tyr1)]Leu-enkephalin
was solved by using 2D NMR spectroscopic techniques.
Binding
of Leu-enkephalin and [RhIII(η5-Cp*)(η6-Tyr1)]Leu-enkephalin
to the recently published crystal structures of the μ- and δ-opioid
receptor is studied. Docking of free Leu-enkephalin reveals two
preferred conformations, one of which suggests an alternative binding
site for the tyrosine residue. Furthermore, the three-dimensional
solution structure of [RhIII(η5-Cp*)(η6-Tyr1)]Leu-enkephalin
was solved by using 2D NMR spectroscopic techniques.
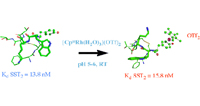 The bioconjugation of organometallic complexes with peptides has proven to be a novel approach for drug discovery. We report the facile and chemoselective reaction of tyrosine-containing G-protein-coupled receptor (GPCR) peptides with [Cp*Rh(H2O)3](OTf)2, in water, at room temperature, and at pH 5–6. We have focused on three important GPCR peptides; namely, [Tyr1]-leu-enkephalin, [Tyr4]-neurotensin(8-13), and [Tyr3]-octreotide, each of which has a different position for the tyrosine residue, together with competing functionalities. Importantly, all other functional groups present, i.e., amino, carboxyl, disulfide, phenyl, and indole, were not prominent sites of reactivity by the Cp*Rh tris aqua complex. Furthermore, the influence of the Cp*Rh moiety on the structure of [Tyr3]-octreotide was characterized by 2D NMR, resulting in the first representative structure of an organometallic-peptide complex. The biological consequences of these Cp*Rh-peptide complexes, with respect to GPCR binding and growth inhibition of MCF7 and HT29 cancer cells, will be presented for [(η6-Cp*Rh-Tyr1)-leu-enkephalin](OTf)2 and [(η6-Cp*Rh-Tyr3)-octreotide](OTf)2.
The bioconjugation of organometallic complexes with peptides has proven to be a novel approach for drug discovery. We report the facile and chemoselective reaction of tyrosine-containing G-protein-coupled receptor (GPCR) peptides with [Cp*Rh(H2O)3](OTf)2, in water, at room temperature, and at pH 5–6. We have focused on three important GPCR peptides; namely, [Tyr1]-leu-enkephalin, [Tyr4]-neurotensin(8-13), and [Tyr3]-octreotide, each of which has a different position for the tyrosine residue, together with competing functionalities. Importantly, all other functional groups present, i.e., amino, carboxyl, disulfide, phenyl, and indole, were not prominent sites of reactivity by the Cp*Rh tris aqua complex. Furthermore, the influence of the Cp*Rh moiety on the structure of [Tyr3]-octreotide was characterized by 2D NMR, resulting in the first representative structure of an organometallic-peptide complex. The biological consequences of these Cp*Rh-peptide complexes, with respect to GPCR binding and growth inhibition of MCF7 and HT29 cancer cells, will be presented for [(η6-Cp*Rh-Tyr1)-leu-enkephalin](OTf)2 and [(η6-Cp*Rh-Tyr3)-octreotide](OTf)2.
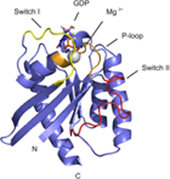 Rheb is a homolog of Ras GTPase that regulates cell growth, prolife
ration, and regeneration via mammalian target of rapamycin (mTOR). Because of the well established potential of activated Ras to promote survival, we sought to investigate the ability of Rheb signaling to phenocopy Ras. We found that overexpression of lipid-anchored Rheb enhanced the apoptotic effects induced by UV light, TNFα, or tunicamycin in an mTOR complex 1 (mTORC1)-dependent manner. Knocking down endogenous Rheb or applying rapamycin led to partial protection, identifying Rheb as a mediator of cell death. Ras and c-Raf kinase opposed the apoptotic effects induced by UV light or TNFα but did not prevent Rheb-mediated apoptosis. To gain structural insight into the signaling mechanisms, we determined the structure of Rheb-GDP by NMR. The complex adopts the typical canonical fold of RasGTPases and displays the characteristic GDP-dependent picosecond to nanosecond backbone dynamics of the switch I and switch II regions. NMR revealed Ras effector-like binding of activated Rheb to the c-Raf-Ras-binding domain (RBD), but the affinity was 1000-fold lower than the Ras/RBD interaction, suggesting a lack of functional interaction. shRNA-mediated knockdown of apoptosis signal-regulating kinase 1 (ASK-1) strongly reduced UV or TNFα-induced apoptosis and suppressed enhancement by Rheb overexpression. In conclusion, Rheb-mTOR activation not only promotes normal cell growth but also enhances apoptosis in response to diverse toxic stimuli via an ASK-1-mediated mechanism. Pharmacological regulation of the Rheb/mTORC1 pathway using rapamycin should take the presence of cellular stress into consideration, as this may have clinical implications.
Rheb is a homolog of Ras GTPase that regulates cell growth, prolife
ration, and regeneration via mammalian target of rapamycin (mTOR). Because of the well established potential of activated Ras to promote survival, we sought to investigate the ability of Rheb signaling to phenocopy Ras. We found that overexpression of lipid-anchored Rheb enhanced the apoptotic effects induced by UV light, TNFα, or tunicamycin in an mTOR complex 1 (mTORC1)-dependent manner. Knocking down endogenous Rheb or applying rapamycin led to partial protection, identifying Rheb as a mediator of cell death. Ras and c-Raf kinase opposed the apoptotic effects induced by UV light or TNFα but did not prevent Rheb-mediated apoptosis. To gain structural insight into the signaling mechanisms, we determined the structure of Rheb-GDP by NMR. The complex adopts the typical canonical fold of RasGTPases and displays the characteristic GDP-dependent picosecond to nanosecond backbone dynamics of the switch I and switch II regions. NMR revealed Ras effector-like binding of activated Rheb to the c-Raf-Ras-binding domain (RBD), but the affinity was 1000-fold lower than the Ras/RBD interaction, suggesting a lack of functional interaction. shRNA-mediated knockdown of apoptosis signal-regulating kinase 1 (ASK-1) strongly reduced UV or TNFα-induced apoptosis and suppressed enhancement by Rheb overexpression. In conclusion, Rheb-mTOR activation not only promotes normal cell growth but also enhances apoptosis in response to diverse toxic stimuli via an ASK-1-mediated mechanism. Pharmacological regulation of the Rheb/mTORC1 pathway using rapamycin should take the presence of cellular stress into consideration, as this may have clinical implications.
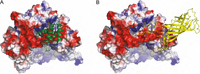 Psb27 is a membrane-extrinsic subunit of photosystem II (PSII) where it is involved in the assembly and maintenance of this large membrane protein complex that catalyzes one of the key reactions in the biosphere, the light-induced oxidation of water. Here, we report for the first time the structure of Psb27 that was not observed in the previous crystal structures of PSII due to its transient binding mode. The Psb27 structure shows that the core of the protein is a right-handed four-helix bundle with an up-down-up-down topology. The electrostatic potential of the surface generated by the amphipathic helices shows a dipolar distribution which fits perfectly to the major PsbO binding site on the PSII complex. Moreover, the presented docking model could explain the function of Psb27 (A), which prevents the binding of PsbO to facilitate the assembly of the Mn4Ca cluster (B).
Psb27 is a membrane-extrinsic subunit of photosystem II (PSII) where it is involved in the assembly and maintenance of this large membrane protein complex that catalyzes one of the key reactions in the biosphere, the light-induced oxidation of water. Here, we report for the first time the structure of Psb27 that was not observed in the previous crystal structures of PSII due to its transient binding mode. The Psb27 structure shows that the core of the protein is a right-handed four-helix bundle with an up-down-up-down topology. The electrostatic potential of the surface generated by the amphipathic helices shows a dipolar distribution which fits perfectly to the major PsbO binding site on the PSII complex. Moreover, the presented docking model could explain the function of Psb27 (A), which prevents the binding of PsbO to facilitate the assembly of the Mn4Ca cluster (B).
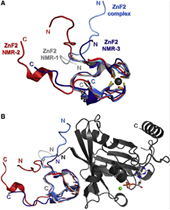 Nucleoporin (Nup) 153 is a highly mobile, multifunctional, and essential nuclear pore protein. It contains four zinc finger motifs that are thought to be crucial for the regulation of transportreceptor/ cargo interactions via their binding to the small guanine nucleotide binding protein, Ran. We found this interaction to be independent of the phosphorylation state of the nucleotide. Ran binds with the highest affinity to the second zinc finger motif of Nup153 (Nup153ZnF2). Here we present the crystal structure of this complex (B), revealing a new type of Ran-Ran interaction partner interface together with the solution structure of Nup153ZnF2 (A). According to our complex structure, Nup153ZnF2 binding to Ran excludes the formation of a Ran-importin-b complex. This finding suggests a local Nup153-mediated Ran reservoir at the nucleoplasmic distal ring of the nuclear pore, where nucleotide exchange may take place in a ternary Nup153-Ran-RCC1 complex, so that import complexes are efficiently terminated.
Nucleoporin (Nup) 153 is a highly mobile, multifunctional, and essential nuclear pore protein. It contains four zinc finger motifs that are thought to be crucial for the regulation of transportreceptor/ cargo interactions via their binding to the small guanine nucleotide binding protein, Ran. We found this interaction to be independent of the phosphorylation state of the nucleotide. Ran binds with the highest affinity to the second zinc finger motif of Nup153 (Nup153ZnF2). Here we present the crystal structure of this complex (B), revealing a new type of Ran-Ran interaction partner interface together with the solution structure of Nup153ZnF2 (A). According to our complex structure, Nup153ZnF2 binding to Ran excludes the formation of a Ran-importin-b complex. This finding suggests a local Nup153-mediated Ran reservoir at the nucleoplasmic distal ring of the nuclear pore, where nucleotide exchange may take place in a ternary Nup153-Ran-RCC1 complex, so that import complexes are efficiently terminated.
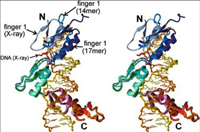 The zinc finger domain of the Wilms tumor suppressor protein (WT1) contains four canonical Cys2His2 zinc fingers. WT1 binds preferentially to DNA sequences that are closely related to the EGR-1 consensus site. We report the structure determination by both X-ray crystallography and NMR spectroscopy of the WT1 zinc finger domain in complex with DNA. The X-ray structure was determined for the complex with a cognate 14 base-pair oligonucleotide, and composite X-ray/NMR structures were determined for complexes with both the 14 base-pair and an extended 17 base-pair DNA. This combined approach allowed unambiguous determination of the position of the first zinc finger, which is influenced by lattice contacts in the crystal structure. The crystal structure shows the second, third and fourth zinc finger domains inserted deep into the major groove of the DNA where they make base-specific interactions. The DNA duplex is distorted in the vicinity of the first zinc finger, with a cytidine twisted and tilted out of the base stack to pack against finger 1 and the tip of finger 2. By contrast, the composite X-ray/NMR structures show that finger 1 continues to follow the major groove in the solution complexes. However, the orientation of the helix is non-canonical, and the fingertip and the N terminus of the helix project out of the major groove; as a consequence, the zinc finger side-chains that are commonly involved in base recognition make no contact with the DNA. We conclude that finger 1 helps to anchor WT1 to the DNA by amplifying the binding affinity although it does not contribute significantly to binding specificity. The structures provide molecular level insights into the potential consequences of mutations in zinc fingers 2 and 3 that are associated with Denys-Drash syndrome and nephritic syndrome. The mutations are of two types, and either destabilize the zinc finger structure or replace key base contact residues.
The zinc finger domain of the Wilms tumor suppressor protein (WT1) contains four canonical Cys2His2 zinc fingers. WT1 binds preferentially to DNA sequences that are closely related to the EGR-1 consensus site. We report the structure determination by both X-ray crystallography and NMR spectroscopy of the WT1 zinc finger domain in complex with DNA. The X-ray structure was determined for the complex with a cognate 14 base-pair oligonucleotide, and composite X-ray/NMR structures were determined for complexes with both the 14 base-pair and an extended 17 base-pair DNA. This combined approach allowed unambiguous determination of the position of the first zinc finger, which is influenced by lattice contacts in the crystal structure. The crystal structure shows the second, third and fourth zinc finger domains inserted deep into the major groove of the DNA where they make base-specific interactions. The DNA duplex is distorted in the vicinity of the first zinc finger, with a cytidine twisted and tilted out of the base stack to pack against finger 1 and the tip of finger 2. By contrast, the composite X-ray/NMR structures show that finger 1 continues to follow the major groove in the solution complexes. However, the orientation of the helix is non-canonical, and the fingertip and the N terminus of the helix project out of the major groove; as a consequence, the zinc finger side-chains that are commonly involved in base recognition make no contact with the DNA. We conclude that finger 1 helps to anchor WT1 to the DNA by amplifying the binding affinity although it does not contribute significantly to binding specificity. The structures provide molecular level insights into the potential consequences of mutations in zinc fingers 2 and 3 that are associated with Denys-Drash syndrome and nephritic syndrome. The mutations are of two types, and either destabilize the zinc finger structure or replace key base contact residues.
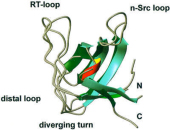 Melanoma inhibitory activity (MIA) protein is a clinically valuable marker in patients with malignant melanoma, as enhanced values diagnose metastatic melanoma stages III and IV. Here we show that the recombinant human MIA adopts an SH3 domain-like fold in solution, with two perpendicular, antiparallel, three- and five-stranded ß-sheets. In contrast to known structures with the SH3 domain fold, MIA is a single-domain protein, and contains an additional antiparallel ß-sheet and two disulfide bonds. MIA is also the first extracellular protein found to have the SH3 domain-like fold. Furthermore, we show that MIA interacts with fibronectin and that the peptide ligands identified for MIA exhibit a matching sequence to type III human fibronectin repeats, especially to FN14, which is close to an integrin alpha4beta1 binding site. The present study, therefore, may explain the role of MIA in metastasis in vivo, and supports a model in which the binding of human MIA to type III repeats of fibronectin competes with integrin binding, thus detaching cells from the extracellular matrix.
Melanoma inhibitory activity (MIA) protein is a clinically valuable marker in patients with malignant melanoma, as enhanced values diagnose metastatic melanoma stages III and IV. Here we show that the recombinant human MIA adopts an SH3 domain-like fold in solution, with two perpendicular, antiparallel, three- and five-stranded ß-sheets. In contrast to known structures with the SH3 domain fold, MIA is a single-domain protein, and contains an additional antiparallel ß-sheet and two disulfide bonds. MIA is also the first extracellular protein found to have the SH3 domain-like fold. Furthermore, we show that MIA interacts with fibronectin and that the peptide ligands identified for MIA exhibit a matching sequence to type III human fibronectin repeats, especially to FN14, which is close to an integrin alpha4beta1 binding site. The present study, therefore, may explain the role of MIA in metastasis in vivo, and supports a model in which the binding of human MIA to type III repeats of fibronectin competes with integrin binding, thus detaching cells from the extracellular matrix.
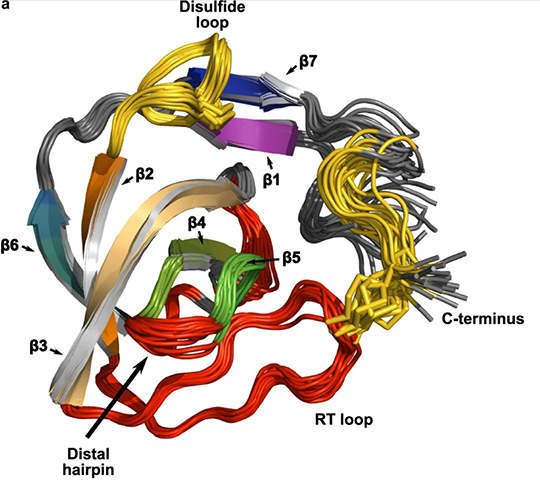
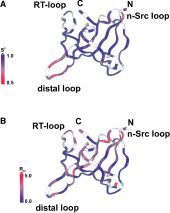 The melanoma inhibitory activity (MIA) protein is a clinically valuable marker in patients with malignant melanoma as enhanced values diagnose metastatic melanoma stages III and IV. Here, we report the backbone dynamics of human MIA studied by 15N NMR relaxation experiments. The folded core of human MIA is found to be rigid, but several loops connecting ß-sheets, such as the RT-loop for example, display increased mobility on picosecond to nanosecond time scales (A). One of the most important dynamic features is the pronounced flexibility of the distal loop, comprising residues Asp 68 to Ala 75, where motions on time scales up to milliseconds occur. Further, significant exchange contributions are observed for residues of the canonical binding site of SH3 domains including the RT-loop, the n-Src loop, for the loop comprising residues 13 to 19, which we refer to as the"disulfide loop", in part for the distal loop, and the carboxyl terminus of human MIA (B). The functional importance of this dynamic behavior is discussed with respect to the biological activity of several point mutations of human MIA. The results of this study suggest that the MIA protein and the recently identified highly homologous fibrocyte-derived protein (FDP)/MIA-like (MIAL) constitute a new family of secreted proteins that adopt an SH3 domain-like fold in solution with expanded ligand interactions.
The melanoma inhibitory activity (MIA) protein is a clinically valuable marker in patients with malignant melanoma as enhanced values diagnose metastatic melanoma stages III and IV. Here, we report the backbone dynamics of human MIA studied by 15N NMR relaxation experiments. The folded core of human MIA is found to be rigid, but several loops connecting ß-sheets, such as the RT-loop for example, display increased mobility on picosecond to nanosecond time scales (A). One of the most important dynamic features is the pronounced flexibility of the distal loop, comprising residues Asp 68 to Ala 75, where motions on time scales up to milliseconds occur. Further, significant exchange contributions are observed for residues of the canonical binding site of SH3 domains including the RT-loop, the n-Src loop, for the loop comprising residues 13 to 19, which we refer to as the"disulfide loop", in part for the distal loop, and the carboxyl terminus of human MIA (B). The functional importance of this dynamic behavior is discussed with respect to the biological activity of several point mutations of human MIA. The results of this study suggest that the MIA protein and the recently identified highly homologous fibrocyte-derived protein (FDP)/MIA-like (MIAL) constitute a new family of secreted proteins that adopt an SH3 domain-like fold in solution with expanded ligand interactions.
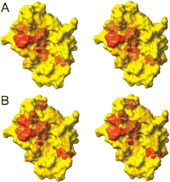 The oncoprotein MDM2 inhibits the tumor suppressor protein p53 by binding to the p53 transactivation domain. The p53 gene is inactivated in many human tumors either by mutations or by binding to oncogenic proteins. In some tumors, such as soft tissue sarcomas, overexpression of MDM2 inactivates an otherwise intact p53, disabling the genome integrity checkpoint and allowing cell cycle progression of defective cells. Disruption of the MDM2/p53 interaction leads to increased p53 levels and restored p53 transcriptional activity, indicating restoration of the genome integrity check and therapeutic potential for MDM2/p53 binding antagonists. Here, we show by multidimensional NMR spectroscopy that chalcones (1,3-diphenyl-2-propen-1-ones) are MDM2 inhibitors that bind to a subsite of the p53 binding cleft of human MDM2 (A and B). Biochemical experiments showed that these compounds can disrupt the MDM2/p53 protein complex, releasing p53 from both the p53/MDM2 and DNA-bound p53/MDM2 complexes. These results thus offer a starting basis for structure-based drug design of cancer therapeutics.
The oncoprotein MDM2 inhibits the tumor suppressor protein p53 by binding to the p53 transactivation domain. The p53 gene is inactivated in many human tumors either by mutations or by binding to oncogenic proteins. In some tumors, such as soft tissue sarcomas, overexpression of MDM2 inactivates an otherwise intact p53, disabling the genome integrity checkpoint and allowing cell cycle progression of defective cells. Disruption of the MDM2/p53 interaction leads to increased p53 levels and restored p53 transcriptional activity, indicating restoration of the genome integrity check and therapeutic potential for MDM2/p53 binding antagonists. Here, we show by multidimensional NMR spectroscopy that chalcones (1,3-diphenyl-2-propen-1-ones) are MDM2 inhibitors that bind to a subsite of the p53 binding cleft of human MDM2 (A and B). Biochemical experiments showed that these compounds can disrupt the MDM2/p53 protein complex, releasing p53 from both the p53/MDM2 and DNA-bound p53/MDM2 complexes. These results thus offer a starting basis for structure-based drug design of cancer therapeutics.
 The conformation of thymosin beta 9 in solution of 40% (v/v) 1,1,1,3,3,3-hexafluoro-2-propanol-d2 in water has been investigated by two-dimensional 1H-nmr spectroscopy. Under this condition thymosin beta 9 adopts an ordered structure. The determination of the conformation of the peptide was based on a set of 304 approximate interproton distance constraints derived from nuclear Overhauser enhancement measurements. The conformation of thymosin beta 9 includes two helical regions from residues 4 to 27 and 32 to 41. The two helices are separated by a poorly defined loop region between amino acids 28 and 31; the N-terminus of thymosin beta 9 shows random-coil structure only.
The conformation of thymosin beta 9 in solution of 40% (v/v) 1,1,1,3,3,3-hexafluoro-2-propanol-d2 in water has been investigated by two-dimensional 1H-nmr spectroscopy. Under this condition thymosin beta 9 adopts an ordered structure. The determination of the conformation of the peptide was based on a set of 304 approximate interproton distance constraints derived from nuclear Overhauser enhancement measurements. The conformation of thymosin beta 9 includes two helical regions from residues 4 to 27 and 32 to 41. The two helices are separated by a poorly defined loop region between amino acids 28 and 31; the N-terminus of thymosin beta 9 shows random-coil structure only.