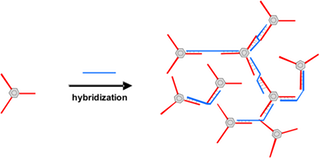
DNA side chain polymers will be prepared by statistical copolymerization of ss DNA macromonomers.(D. C. Lin, B. Yurke, N. A. Langrana, Journal of Biomechanical Engineering-Transactions of the ASME 2004, 126, 104.) An oligonucleotide (oDNA) is functionalized at one end with a polymerizable unit that is subsequently transformed into a side chain architecture by polymerization with a conventional monomer, in this case acrylamide derivatives. These DNA side chain polymers can be transformed into cross linked structures by two different methods. Either two of these conjugates with complementary sequences are hybridized for network formation. The second possibility makes use of cross linking DNA (Figure A).

The DNA side chain polymer is mixed
with an oDNA that encodes two times the complementary sequence of the nucleic
acid/polymer hybrid.(S. Nagahara, T. Matsuda, Polymer Gels and
Networks 1996, 4, 111.) An appealing aspect of these structures is that spatially well
defined hydrogels can be generated.
For the preparation of all DNA networks instead of DNA side chain polymers we will employ DNA star polymers (Figure B). Characteristic for such structures is that at least three oDNAs are covalently connected to a core molecule.
 Their synthesis will be carried out by reacting a
tris-phosphoramidite containing a
1,3,5-functionalized benzene derivative with three oDNAs that are still present
on the solid support. In a similar way triblock copolymers have been generated
where two oDNAs were coupled to a bifunctional polymer unit.(Alemdaroglu FE, Safak M, Wang J, Berger R, Herrmann A, Chem. Commun. 13: 1358-1359 2007) Crosslinking can then be achieved in a similar way
as described for the side chain polymer structures. Especially appealing in
regard to networks based on three-arm polymer structures is that
compartmentalization can be coupled to replication since trisoligonucleotides
were successfully build up from template molecules allowing successfully
chemical copying of connectivity.(Eckardt LH, Naumann K, Pankau WM, Rein M, Schweitzer M, Windhab N, von Kiedrowski G Nature 420 (6913): 286-286
2002)
Their synthesis will be carried out by reacting a
tris-phosphoramidite containing a
1,3,5-functionalized benzene derivative with three oDNAs that are still present
on the solid support. In a similar way triblock copolymers have been generated
where two oDNAs were coupled to a bifunctional polymer unit.(Alemdaroglu FE, Safak M, Wang J, Berger R, Herrmann A, Chem. Commun. 13: 1358-1359 2007) Crosslinking can then be achieved in a similar way
as described for the side chain polymer structures. Especially appealing in
regard to networks based on three-arm polymer structures is that
compartmentalization can be coupled to replication since trisoligonucleotides
were successfully build up from template molecules allowing successfully
chemical copying of connectivity.(Eckardt LH, Naumann K, Pankau WM, Rein M, Schweitzer M, Windhab N, von Kiedrowski G Nature 420 (6913): 286-286
2002)The simplest way to electronically disassemble both kinds of networks is to raise the pH by electrolysis of water at electrodes within the microfluidic system.
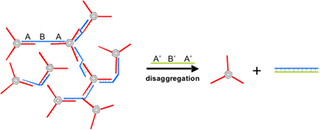
Another way to reverse the network structure consists of employing larger cross-linking strands that remain partially unhybridized during network formation. When now a full complement of the cross-linking strand (a so called removal strand) is added, the network disassociates due to the greater duplex stability of the cross-linking and the removal strand (Figure C). The basis for this switching process relies on DNA sequences that have been developed in the context of DNA-fuelled machines. (B. Yurke, A. J. Turberfield, A. P. Mills, F. C. Simmel, J. L. Neumann, Nature 2000, 406, 605.; L. P. Feng, S. H. Park, J. H. Reif, H. Yan, Angew. Chem. Int. Ed. 2003, 42, 4342.)
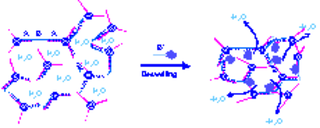
The concept of programmable gelation can now further be extended to combine cross-linking with deswelling of the DNA hydrogel using the latter architecture with unhybridized cross-links. When the ss region of the crosslinking strand is hybridized with an amphiphilic DNA diblock copolymer, e.g DNA-b-PPO, deswelling is induced due to the presence of the hydrophobic synthetic polymer segment (Figure D). The contraction process is due to repelling water out of the hydrogel’s pores. In this way networks responding to multiple stimuli will be obtained and state of gelation will be fully controllable by computational input signals.
